Master of Science (MSc), Doctor of Philosophy (PhD)
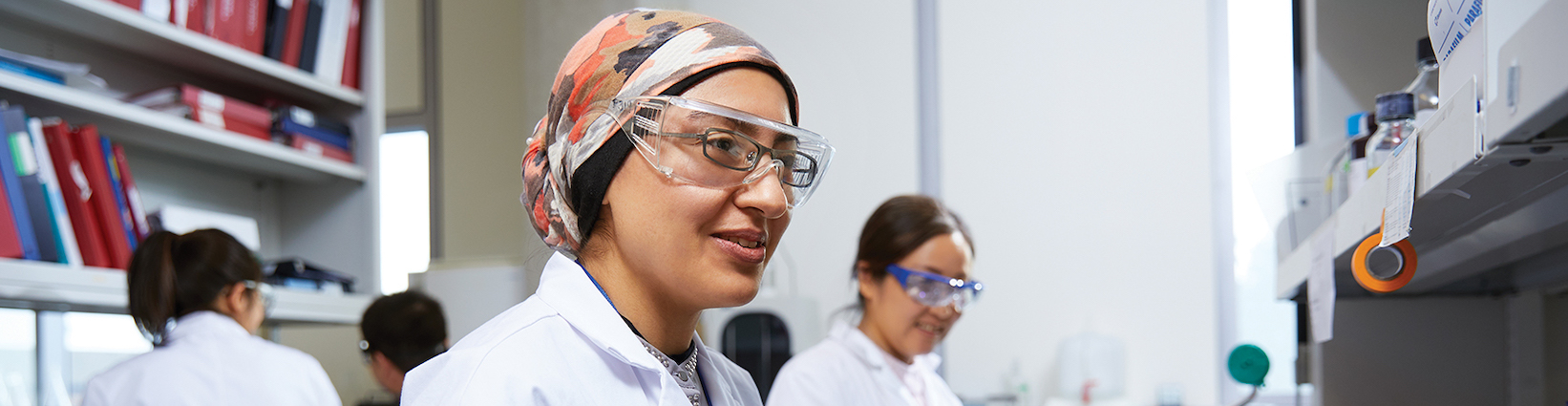
As one of the largest Biology departments in Canada, we produce cutting-edge, hands-on research and advance the current frontiers of knowledge guided by award-winning and internationally recognized faculty and researchers (e.g. Steacie Fellows, Young Investigator Award, York Research Chairs, Canada Research Chairs).
Biology offers graduate programs leading to the Master of Science degree (by research thesis) and the Doctor of Philosophy degree (by research dissertation). We do not offer degrees based solely on coursework. Prospective students must obtain a commitment from a faculty supervisor prior to being accepted into the program.
Both the M.Sc. and Ph.D. are designed to give students in-depth knowledge of a specific area of current biology. This is accomplished primarily by hands-on research on an important problem or question that advances the current frontiers of knowledge and most of our students publish their thesis research in scientific journals.
Areas of Specialization
Cell and Molecular Biology
Ecology and Evolution
Physiology and Neuroscience
Featured Faculty
Amanda Liczner, PhD '20
How important are bumble bees in our society? As a recent graduate of the Biology graduate program and postdoctoral fellow at the UBC's Okanagan campus, Liczner discusses her research with us, which examines the habitat characteristics of bumble bees in North America and their significance to the agricultural sector.

Learn More
The Graduate Program in Biology at York is an exciting environment to pursue innovative, socially engaging, career-ready education. Contact our Graduate Program Assistant to learn more.